We use cookies to enhance your browsing experience and analyze site traffic. By continuing to use this site, you agree to our cookie policy. Privacy Policy
Epitope Prediction
Our Approach
Provide antigen structure and antibody sequence, and the INSPIRE-SEEN model will predict structural epitopes with extremely high accuracy, providing critical insights for antibody screening and differentiated design. 100x lower cost than traditional experiments, 28% accuracy improvement over existing computational methods, with results in about 10-20 minutes.
Validated Results
Why Is This Needed?
"Epitopes are directly related to antibody function, mechanism of action, safety, and efficacy. The binding epitope determines molecular function, clinical indications, and patent landscape. Early identification and evaluation of potential binding epitopes is critical for optimizing efficacy and reducing homogeneity risks."
Key Advantages
Key Advantages
Long Cycles, High Costs
Experimental approaches typically take 4-8 weeks with costs ranging from tens to hundreds of thousands per experiment; traditional high-throughput screening and structural analysis are time-consuming.
Expression Difficulty
Some proteins and antibodies are extremely difficult to express, making it hard to obtain epitope information via traditional experiments, especially for complex membrane protein targets.
Inaccurate Computational Predictions
Existing computational methods like AF2-multimer and HADDOCK still have insufficient accuracy (~50%) for predicting antibody-antigen binding modes.
Homogeneity & Resistance Bottlenecks
High overlap with existing antibody epitopes means similar competitive mechanisms; tumor cells escape via epitope mutations — lacking differentiated epitopes makes it hard to break through barriers.
Highly Accurate, Continuously Evolving
Prediction accuracy improved 28% over AF2-multimer (from 50% to 78%), continuously trained with massive internal cryo-EM data.
Ultra-Fast & Low Cost
Results in about 10-20 minutes from submission, 100x to 1000x cheaper than traditional wet lab experiments.
Accelerate Differentiated Design
Help R&D teams avoid known epitopes, lock in new epitope ranges early; high-scoring epitopes can be used for targeted library screening or cell-based validation.
Validation
Case Studies
In a test set of 110 known epitopes, INSPIRE-SEEN demonstrated capabilities exceeding current mainstream AI models. Using HER2 as a representative case to validate the agreement between predicted and actual binding epitopes.
Patch binary classification accuracy of 78%, top-scoring patch accuracy significantly exceeding competitors
On the standard test set, INSPIRE-SEEN achieved 78% Patch binary classification accuracy, far exceeding AF2-multimer's 50%.
| Model | Recall | Precision | AUROC | AUPRC |
|---|---|---|---|---|
| AF2-multimer | 0.40 | 0.43 | 0.65 | 0.48 |
| Internal Model 1 | 0.42 | 0.52 | 0.72 | 0.55 |
| Internal Model 2 | 0.27 | 0.34 | 0.58 | 0.37 |
| INSPIRE-SEEN | 0.52 | 0.55 | 0.71 | 0.60 |
| Model | Rate |
|---|---|
| AF2-multimer | 50% |
| Internal Model 1 | 60% |
| Internal Model 2 | 43% |
| INSPIRE-SEEN | 78% |
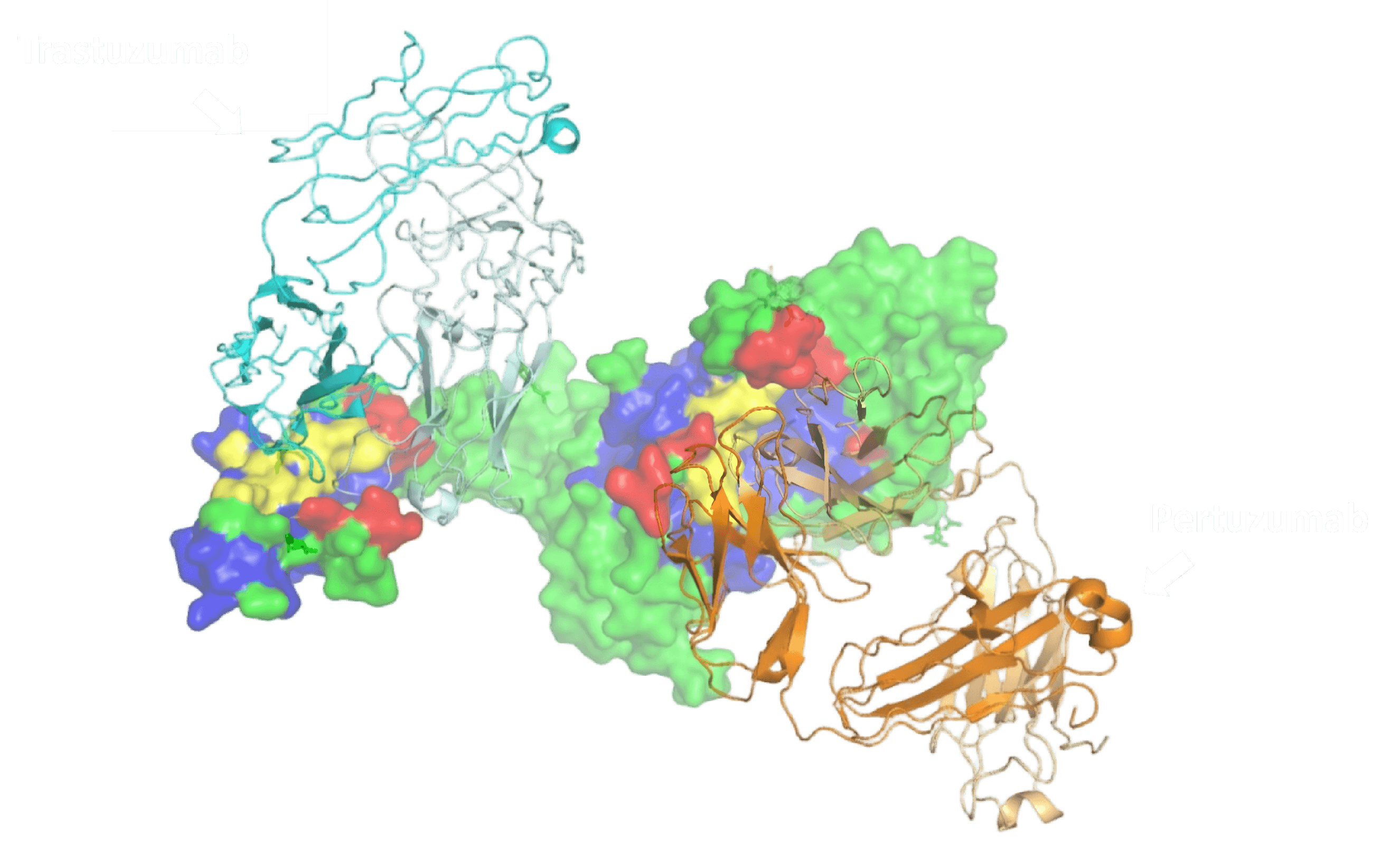
AI model predicted epitopes highly overlap with actual binding epitopes, correctly distinguishing two antibodies' different binding positions
Given HER2 antigen crystal structure and Trastuzumab/Pertuzumab antibody sequences, INSPIRE-SEEN performed deep learning analysis on structural and sequence features, combined with known antibody-antigen interaction data, outputting high-confidence potential epitope predictions. Results show predicted epitope patches overlap with actual binding epitopes and correctly distinguish the two antibodies' different binding positions.
Pipeline
Pipeline
Epitope prediction based on affinity prediction AI models, accurately identifying key amino acid residues by partitioning the antigen surface into multiple patch candidates and calculating their affinity with the antibody.
The system first partitions the antigen surface into patches, then ranks them with an affinity model, and finally outputs the key epitope residues most likely to participate in binding.
Use Cases
Use Cases and Deliverables
Best suited for projects with antibody sequences but unclear likely binding regions on the antigen surface.
Inputs to Prepare
What You Receive
What is the advantage over general complex-prediction models?
Epitope prediction is not just complex pose prediction. It divides the antigen surface into patches and scores binding likelihood. The model is continuously trained with internal cryo-EM complex data and outperforms AF2-multimer on known-epitope tests, making it better suited for antibody epitope localization.
How does it help differentiated antibody design?
By comparing high-scoring patches across candidate antibodies, teams can judge whether candidates avoid known epitopes, approach target functional regions, and provide differentiated epitope space among antibodies.
Accelerate Your Antibody Epitope Identification
The INSPIRE-SEEN model is now open for use. Submit antigen PDB and antibody sequences to get in-depth prediction analysis results within 15 minutes.